By Rob Lewis, CEO, Radical Imaging
Introduction
The Open Health Imaging Foundation (OHIF) platform is now in its third iteration, which is nearing completion and will soon be available as a stable version. This update adds a number of new features for application developers that can greatly improve the overall image viewer experience. This post will talk about the top three new features available in OHIF v3.
The Many Faces of OHIF
Most viewers are pretty static applications; they look how they look and do what they do, but don’t expect them to change any of that when presented with new data or use cases. For that, you’d typically need to launch a whole different application, which can be quite inconvenient.
OHIF v3 behaves essentially like multiple applications in one. This means that you can have different workflows (called “modes”), each focusing on a certain use case, all accessible within one running instance of the OHIF viewer.
In your study list, there could be imaging studies of all different kinds, each requiring very different handling. Perhaps the data formats are different, requiring special decoding or transformation when loading the images (i.e. they need a custom image loader). Maybe the images themselves should be rendered in a particular way requiring a totally different rendering engine. consider how pathology images require different rendering techniques than CT images. A PET/CT study would require yet again a very different sort of layout and rendering. Also consider that each study type would require and benefit from manipulation tools specific to that use case. You wouldn’t measure things in a PET/CT study the same way you would in an Ultrasound study.
How does OHIF do this? It registers each separate workflow at its own route within your application. Each of these routes encapsulates the specific set of image loaders, layouts, rendering engines and tools that make sense for that kind of image data. For instance, you can have a Microscopy workflow and tools at `/microscopy` while a completely different use case of Total Metabolic Tumor Volume workflow can be accessed at `/tmtv`. Then, when expanding a study in the study list, all available modes show up as buttons where only the modes applicable to the selected study show as active and clickable.
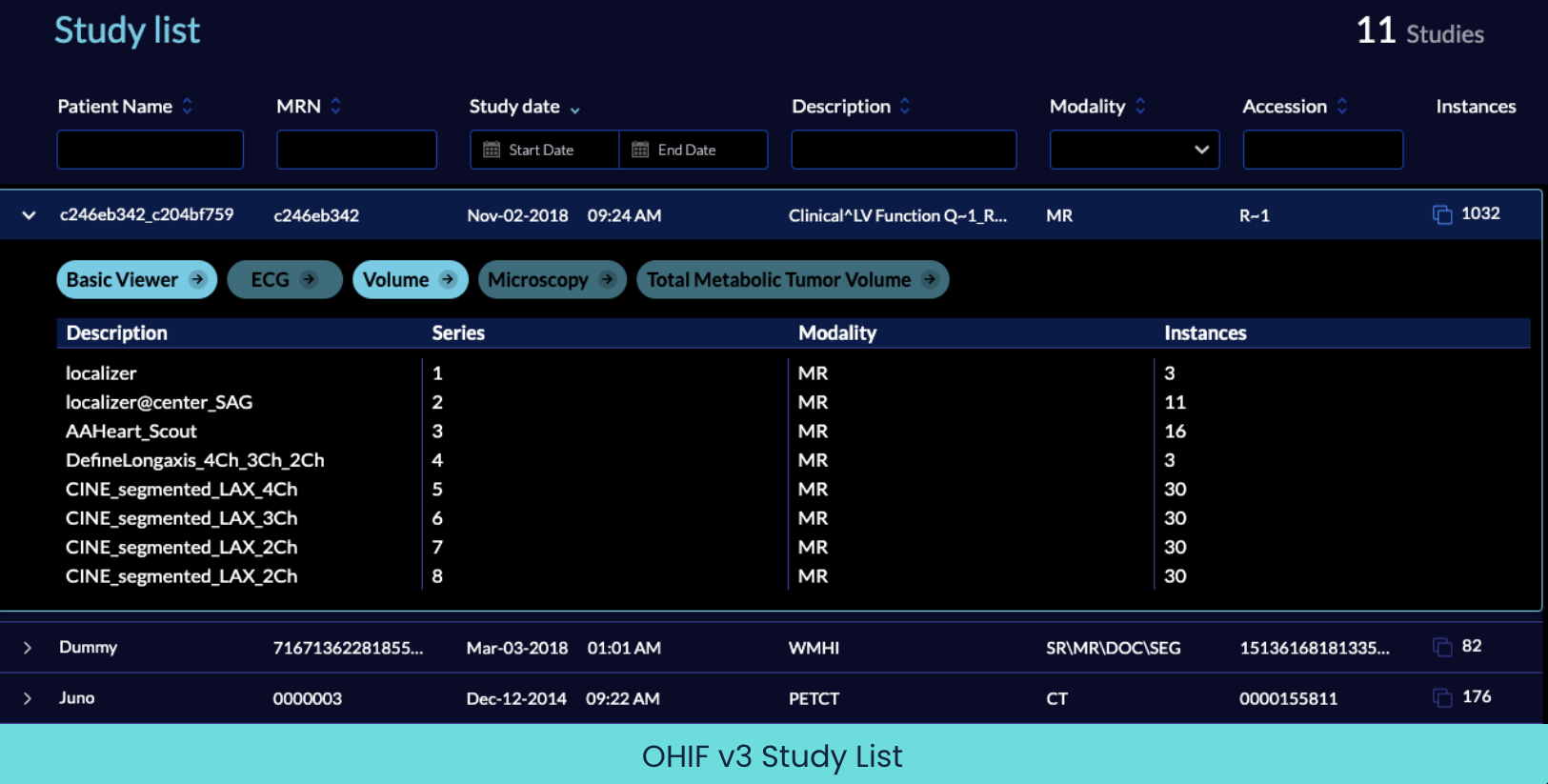
A viewer is useless on its own; you need to be able to get images into it, work with them, then export outputs like modified images, measurements, reports, and such. OHIF v3 is more flexible in the way it can exchange data with other applications, which amplifies its ability to be a useful bridge between applications.
Now you can communicate to various data sources within the same running OHIF instance. For example, you can load images from a DICOMWeb endpoint and export the annotations and results to a different endpoint, like your custom backend. Soon, you will even be able to aggregate multiple image data sources into your single study list, for example from several DICOM nodes, each containing studies you want to look at in OHIF.
OHIF v3 also adds a new data source API, allowing you to specify how to get the metadata separately from the image data. This can be quite useful when they are stored separately, like in many research use cases. You can use this API to write your own custom data source. Here is an example of what it might look like:

You can find more examples here:
Cornercube?
As of v3, OHIF has incorporated Cornerstone3D as its default rendering engine. This is a huge step forward in its visualization capabilities. In the past, there was no easy way to share a volume in GPU memory across viewports, so the same image stack was copied multiple times, reducing performance and eating up tons of that scarce resource. In addition, it was super awkward to coordinate interactions between annotation workflows in 2D and 3D viewports, if it was possible at all.
Now, there is a single offscreen VTK.js viewport which holds the stack in GPU memory, performs all of the rendering (2D and 3D), and moves the result to each of the viewports in the layout. This allows for fast rendering with many viewports and can support more and/or larger stacks. In addition, Cornerstone3D adds support for 3D annotations and 3D segmentation rendering and editing tools, all of which natively work in 3D, showing and editing the overlays in whatever plane is displayed without any extra effort.
Cornerstone3D greatly improves the usability of OHIF-based viewers for complex clinical use cases that require advanced visualizations (e.g. MIP, MPR, etc) and analysis.
You can play with a live PET/CT example here.
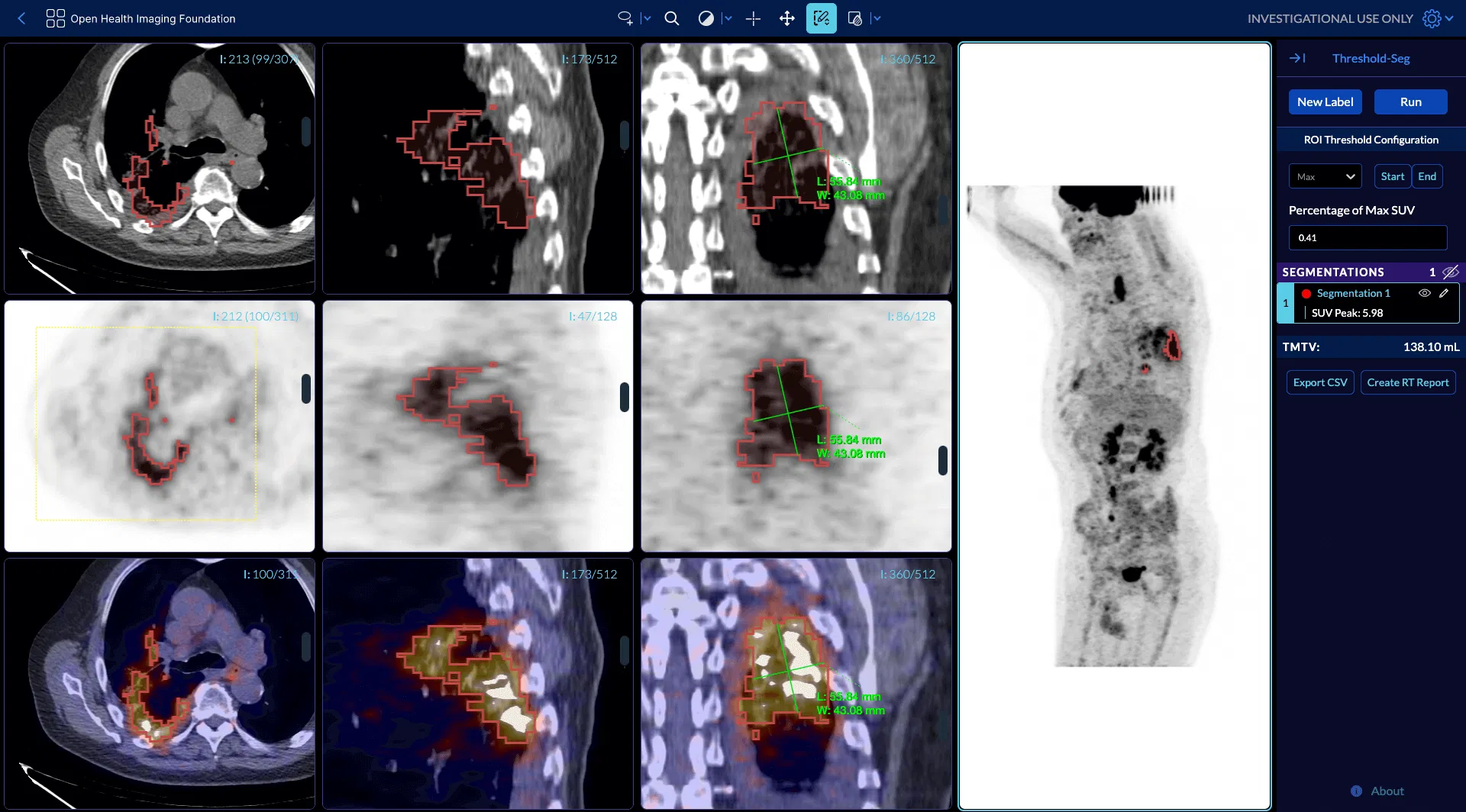
OHIF v2 to OHIF v3 Software Migration
Are you convinced you should migrate to OHIF v3? We can help you sort out what that would take, and even assist with the implementation. Learn more about Radical’s v3 migration service here.
Publish Date
Publish Date
By Rob Lewis, CEO, Radical Imaging
Introduction
The Open Health Imaging Foundation (OHIF) platform is now in its third iteration, which is nearing completion and will soon be available as a stable version. This update adds a number of new features for application developers that can greatly improve the overall image viewer experience. This post will talk about the top three new features available in OHIF v3.
The Many Faces of OHIF
Most viewers are pretty static applications; they look how they look and do what they do, but don’t expect them to change any of that when presented with new data or use cases. For that, you’d typically need to launch a whole different application, which can be quite inconvenient.
OHIF v3 behaves essentially like multiple applications in one. This means that you can have different workflows (called “modes”), each focusing on a certain use case, all accessible within one running instance of the OHIF viewer.
In your study list, there could be imaging studies of all different kinds, each requiring very different handling. Perhaps the data formats are different, requiring special decoding or transformation when loading the images (i.e. they need a custom image loader). Maybe the images themselves should be rendered in a particular way requiring a totally different rendering engine. consider how pathology images require different rendering techniques than CT images. A PET/CT study would require yet again a very different sort of layout and rendering. Also consider that each study type would require and benefit from manipulation tools specific to that use case. You wouldn’t measure things in a PET/CT study the same way you would in an Ultrasound study.
How does OHIF do this? It registers each separate workflow at its own route within your application. Each of these routes encapsulates the specific set of image loaders, layouts, rendering engines and tools that make sense for that kind of image data. For instance, you can have a Microscopy workflow and tools at `/microscopy` while a completely different use case of Total Metabolic Tumor Volume workflow can be accessed at `/tmtv`. Then, when expanding a study in the study list, all available modes show up as buttons where only the modes applicable to the selected study show as active and clickable.

A viewer is useless on its own; you need to be able to get images into it, work with them, then export outputs like modified images, measurements, reports, and such. OHIF v3 is more flexible in the way it can exchange data with other applications, which amplifies its ability to be a useful bridge between applications.
Now you can communicate to various data sources within the same running OHIF instance. For example, you can load images from a DICOMWeb endpoint and export the annotations and results to a different endpoint, like your custom backend. Soon, you will even be able to aggregate multiple image data sources into your single study list, for example from several DICOM nodes, each containing studies you want to look at in OHIF.
OHIF v3 also adds a new data source API, allowing you to specify how to get the metadata separately from the image data. This can be quite useful when they are stored separately, like in many research use cases. You can use this API to write your own custom data source. Here is an example of what it might look like:

You can find more examples here:
Cornercube?
As of v3, OHIF has incorporated Cornerstone3D as its default rendering engine. This is a huge step forward in its visualization capabilities. In the past, there was no easy way to share a volume in GPU memory across viewports, so the same image stack was copied multiple times, reducing performance and eating up tons of that scarce resource. In addition, it was super awkward to coordinate interactions between annotation workflows in 2D and 3D viewports, if it was possible at all.
Now, there is a single offscreen VTK.js viewport which holds the stack in GPU memory, performs all of the rendering (2D and 3D), and moves the result to each of the viewports in the layout. This allows for fast rendering with many viewports and can support more and/or larger stacks. In addition, Cornerstone3D adds support for 3D annotations and 3D segmentation rendering and editing tools, all of which natively work in 3D, showing and editing the overlays in whatever plane is displayed without any extra effort.
Cornerstone3D greatly improves the usability of OHIF-based viewers for complex clinical use cases that require advanced visualizations (e.g. MIP, MPR, etc) and analysis.
You can play with a live PET/CT example here.

OHIF v2 to OHIF v3 Software Migration
Are you convinced you should migrate to OHIF v3? We can help you sort out what that would take, and even assist with the implementation. Learn more about Radical’s v3 migration service here.